Nomenclature of Electron-Transfer Proteins
Contents of this section
Most but not all of the proteins in this group are iron-sulfur proteins. Recommendations for the nomenclature of iron-sulfur proteins were first formulated in 1971 and revised in 1978 (ref 16). Since that time, a number of major contributions to the field have reported structure determination by X-ray crystallography, elaboration of a series of model compounds, and the discovery of novel examples of these proteins. On the basis of the new information, it became desirable to adjust the nomenclature of the iron-sulfur proteins and thereby take into account the more precise description of known species while retaining flexibility for future discoveries.
Whenever possible, terms that have gained acceptance have been retained and particular attention has been given to the development of a useful shorthand notation. However, in order to comply with the established rules for the nomenclature of inorganic compounds (ref 17), it was decided in 1978 to make some changes from usage in the biochemical literature that had developed in the early 1970s. (see Note)
Note The group of 'high potential iron-sulfur proteins' has been deleted on the following basis: a) The three-dimensional structure of the active centre of this protein has been found to be essentially the same as that of ferredoxins (ref 18, 19). The magnetic properties differ because the Fe-S clusters may assume three different oxidation levels (ref 18, 20). Thus, an oxidized ferredoxin (diamagnetic) corresponds to the reduced Chromatium vinosum high-potential iron-sulfur protein (diamagnetic); the extreme oxidation levels, i.e., those of reduced ferredoxin and oxidized C. vinosum high potential iron-sulfur protein, have one electron more or one electron less, respectively, than this oxidation level and are paramagnetic. b) After the original study (ref 21) with the Chromatium high potential iron-sulfur protein, it became apparent that other iron-sulfur proteins may have the unusual oxidation level of the Chromatium protein without a high midpoint oxidation-reduction potential (ref 22). Conversely, ferredoxins with the usual oxidation level and magnetic properties may have high midpoint potentials (ref 23). The expression, 'high potential Fe-S protein' (or HiPIP) has been used increasingly in the literature to indicate the ability to form the higher oxidation level of the Chromatium protein. This practice is inappropriate since it uses a term applying to a redox potential for the description of a magnetic property or an oxidation level.Proteins containing iron can be divided into three groups: hemeproteins, iron-sulfur proteins, and other iron-containing proteins (Scheme I). The last group includes ferritin, transferrin and the oxygenases; of these, the oxygenases also have electron-transfer functions. The term 'iron-sulfur proteins' refers only to those proteins in which a non-heme iron is ligated with inorganic sulfur or cysteine sulfur.
The iron-sulfur proteins (abbreviation: Fe-S proteins) fall into two major categories: simple iron-sulfur proteins and complex iron-sulfur proteins. 'Simple' need only be used when the difference from complex iron-sulfur proteins is emphasized. Simple iron-sulfur proteins contain only one or more Fe-S clusters whereas the complex proteins bear such additional active redox centres as flavin, molybdenum or heme. Often EPR spectra are used for classification and identification of Fe-S clusters in proteins.
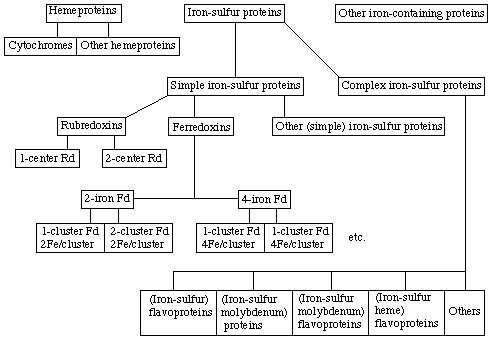
5.2 Types of simple iron-sulfur proteins
Simple iron-sulfur proteins fall into three groups: rubredoxins, ferredoxins and other simple iron-sulfur proteins, as shown in the adjacent scheme I.
Rubredoxins (abbreviation: Rd) comprise those iron-sulfur proteins without acid-labile sulfur that are characterized by having iron in a typical thiolate coordination, i.e. an iron centre surrounded by four cysteine residues or sulfur-containing ligands. More than one iron centre of this type may exist in the molecule. Oxidized rubredoxin usually exhibits the characteristic EPR spectrum of high-spin Fe(III) ion in a rhombic ligand field, g = 4.3; the reduced form gives no discernible EPR signal. Only negative redox potentials at pH 7 have been noted for those rubredoxins presently characterized. Desulforedoxin is a variant rubredoxin with a somewhat higher symmetry and distinctive EPR spectrum with g-values of 7.7, 4.1 and 1.8.
The full name should be listed as follows: (source) rubredoxin, (function), e.g. Pseudomonas oleovorans rubredoxin, alkane w-hydroxylation.
Ferredoxins (abbreviation: Fd) comprise those iron-sulfur proteins that contain more than one iron and labile sulfur atoms and that exclusively display electron-carrier activity but not classical enzyme function. Thus, since hydrogenases (iron-sulfur proteins containing one or more Fe-S clusters) are enzymes, they are not classified as ferredoxins.
The criterion that ferredoxins must have negative oxidation-reduction midpoint potential at pH 7 has been abandoned; there is no need to distinguish between ferredoxins and the previously designated 'high potential iron-sulfur proteins' (see Note with section 5.1) as exemplified by an iron-sulfur protein from Chromatium vinosum (ref 20). Ferredoxins may contain one or more clusters of two or four iron and labile sulfur atoms. In addition to these clusters, more recently both in proteins and in synthetic analogues several other clusters have been reported. (see Note)
Note The existence of Fe-S clusters of novel and unusual stoichiometry has now been proven. The identity of the [3Fe-4S] cluster has been established by chemical analysis, EXAFS and X-ray diffraction (ref 24, 25) methods. In particular there has been a reevaluation of the X-ray structure (ref 26) of the FdI of Azotobacter vinelandii both by the original investigators and by a second group. Both groups concur that the original conclusion of the presence of a 3Fe-3S cluster in this protein was in error and that this cluster is actually of the 3Fe-4S type, comprising a cubane structure with one iron atom missing. Chemical model studies have also indicated that linear structures are possible (ref 25); a linear structure can also be produced in aconitase. When there the Fe/S stoichiometry of a given structure is known, the notation [3Fe-xS] has been introduced; a general form of this notation would be [nFe-xS]. It should be noted that in the absence of precise stoichiometries no formal charge can be assigned to the cluster. Ligands for such clusters are not necessarily cysteine thiolate groups for all metal ions. Evidence has been obtained for some Azotobacter ferredoxins that 3Fe clusters may have a physiological role; the [3Fe-4S] type certainly has a role as intermediate during assembly and degradation.5.2.3 Other iron-sulfur proteins
All simple iron-sulfur proteins that are neither rubredoxins nor ferredoxins will fall into the category of 'other iron-sulfur proteins'. At this point it should be noted that some iron-sulfur proteins are known that do not appear to have a redox role; examples are endonuclease III and aconitase.
5.3 Recommendations for designating iron-sulfur proteins
(a) If any iron-sulfur protein has been given another name previously, this should be stated to minimize confusion.
(b) The term 'high-potential iron-sulfur protein' (abbreviated HiPIP), may continue to be used for the original iron-sulfur protein of photosynthetic bacteria that had been given this name initially. Otherwise, the use of the terms HiPIP or 'high-potential Fe-S protein' is discouraged; there are ferredoxins occurring naturally at the oxidation level of oxidized and reduced spinach ferredoxin that have oxidation-reduction potentials as high as the original HiPIP of bacteria (see Note to section 5.1 ).
(c) At least once in a report, the source of the protein should precede the term rubredoxin, ferredoxin, or iron-sulfur protein. Similarly, for iron-sulfur cluster, or 'centres' of complex iron-sulfur proteins, the proper designation of the parent protein or enzyme should be given, preferably along with the source, e.g. beef-heart NADH dehydrogenase Fe-S cluster 1. Thereafter, the designation "cluster 1" may be used. The proliferation of such other abbreviations as centre N-1, S-1, or bc-1 is discouraged. With regard to more complex systems, including the complex iron-sulfur proteins, it appears to be neither suitable nor desirable to present designations for Fe-S clusters that are inadequately characterized.
(d) Iron-sulfur proteins from the same source that have the same type of Fe-S cluster are numbered sequentially with Roman numerals. A newly isolated iron-sulfur protein that is not fully characterized should be called 'iron-sulfur protein' and given the lowest unused numeral. By analogy, the various iron-sulfur clusters, or 'centres' of an iron-sulfur protein, should be designated Fe-S cluster 1, Fe-S cluster 2, and so on in the order of discovery. Arabic numerals should be used for the different clusters of the same protein or complex, since iron-sulfur proteins and the complexes of the respiratory chain are designated by Roman numerals.
(e) It is useful to present midpoint redox potentials, light absorption and EPR characteristics, particularly when an iron-sulfur protein is first mentioned in a publication.
(a) The designation of a cluster in an iron-sulfur protein containing labile sulfur atoms should consist of square brackets about the number of iron and labile sulfur atoms. Thus, [2Fe-2S] represents a two-iron, two-labile-sulfur cluster and [4Fe-4S], a four-iron, four-labile-sulfur cluster. The protein incorporating such a cluster may be called a 'two-iron-two-sulfur' or 'four-iron-four-sulfur' ferredoxin or iron-sulfur protein. The more recently found [3Fe-4S] clusters, in some cases derived from [4Fe-4S] clusters by loss of Fe, may be called 'three-iron-four-sulfur' ferredoxin or iron-sulfur protein. See also Note to section 5.2.2.
[Comment: The use of hyphens and parentheses (including square brackets) is firmly codified in the nomenclature of inorganic chemistry (ref 17). In order to avoid confusion with established practices in coordination chemistry, a short hyphen is used to write Fe-S and the term is placed between square brackets instead of parentheses.]
(b) The presence of several clusters, as in clostridial ferredoxin, is indicated as follows: 2[4Fe-4S]. This may be called a two-cluster four-iron-four-sulfur ferredoxin or iron-sulfur protein.
(c) When the formal charge of the cluster is calculated, the sulfur atoms of the bound cysteine residues are not included in the calculation, contrary to the widespread practice in the earlier literature; thus, for the oxidized and reduced forms, respectively, we have:
| Spinach ferredoxin | [2Fe-2S]2+:[2Fe-2S]1+ |
| Bacillus polymyxa ferredoxin | [4Fe-4S]2+:[4Fe-4S]1+ |
| Clostridium pasteurianum ferredoxin | 2[4Fe-4S]2+:2[4Fe-4S]1+ |
[Comment: The charges referred to are those within the entire cluster. This may seem in contrast with usage in inorganic chemistry where a charge shown in such a formula often refers to the whole coordination entity, i.e. including the other ligands (ref 17). There is also a possibility for misunderstanding the codified use of the term, oxidation state, which usually characterizes a single atom but never a group of atoms (ref 17). In order to avoid confusion, the term 'oxidation level' is used to refer to the cluster as such, and indicated in the square brackets.]
(d) For ferredoxins at the oxidation level typical of oxidized Chromatium HiPIP, the expression [4Fe-4S]n+ (n = 1,2,3), may differentiate between what was formerly called HiPIP and a ferredoxin.
(e) If the oxidation levels in which a ferredoxin can occur are known, this may be indicated as follows for a [4Fe-4S] ferredoxin that is generally obtained on isolation at the (2+) level: [4Fe-4S]2+(3+:2+:1+). This designation implies that the ferredoxin can occur at all three possible oxidation levels. However, these designations should only be used to refer to oxidation levels that can be reached in a biological milieu, i.e. in the absence of agents denaturing the protein, even though artificial oxidants may be used to attain such oxidation levels. By this designation, the highest oxidation levels normally found in Chromatium, HiPIP and the protein from Bacillus polymyxa, both [4Fe-4S] ferredoxins, may be differentiated:
| Chromatium vinosum HiPIP | [4Fe-4S]2+(3+:2+) |
| Bacillus polymyxa ferredoxin | [4Fe-4S]2+(2+:1+) |
(f) Rubredoxins are treated in an analogous fashion, except that the basic centre is designated [Rd] since there is no ambiguity concerning numbers of metal atoms involved. Rubredoxins with multiple centres are denoted as n[Rd]. The formal charges of rubredoxins are [Rd]3+ and [Rd]2+ for the oxidized and reduced forms, respectively.
The examples given in Table II illustrate and contrast the new designations with those applied previously.
Table II: Designation of iron-sulfur proteins as generally obtained on isolation
| Previous designation | Recommended designation |
| Spinach chloroplast ferredoxin | Spinach chloroplast [2Fe-2S]2+Fd or, in a specific context, [2Fe-2S]2+(2+:1+)Fd |
| Azotobacter vinelandii iron-sulfur protein I | Azotobacter vinelandii [2Fe-2S]2+Fe-S protein I |
| Chromatium vinosum high-potential iron-sulfur protein | Chromatium vinosum [4Fe-4S]2+Fd or, in a specific context, [4Fe-4S]2+(3+:2+)Fd |
| Clostridium pasteurianum ferredoxin | Clostridium pasteurianum 2[4Fe4S]2+Fd or, in a specific context, 2[4Fe-4S]2+(2+:1+)Fd |
Iron-sulfur proteins with clusters of different types may be designated in a similar manner as for the above examples.
It seems appropriate to have a nomenclature system that allows for other clusters. According to model work (ref 27) and Mössbauer spectroscopy on a Fe-S protein (ref 28), it has become clear that changing the terminal ligands of iron atoms in a cluster to groups that do not contain sulfur, does not greatly changes the EPR spectra and other properties. The tacit assumption that the terminal ligands are thiolates therefore no longer seems justified for Fe-S clusters in proteins.
In those cases of ligands other than thiolates, it is needed to have a notation for these systems. In the presently available shorthand notation, e.g. [4Fe-4S], the terminal ligands are not considered and no changes are therefore necessary. We now recommend, however, that, in cases where the terminal ligands are to be specifically defined, the same notation be used as for coordination compounds in general and in particular for chemical models, e.g. [Fe4S4(RS)2(L)2]2- (ref 29) for the diamagnetic state of a [4Fe-4S] cluster with two different sets of terminal ligands; in this case the charge of the terminal ligands (i.e. both for the thiolates and the other ligands), must be included in the cluster charge, outside the square brackets.
[Comment: In biochemistry many authors still use the designation Fe2S2 or Fe4S4 for [2Fe-2S] or [4Fe-4S] clusters, respectively. Within the framework of the nomenclature recommended above, where the notations [2Fe-2S], etc., are merely to serve as convenient shorthand, it would be appropriate to use designations such as Fe2S2 or Fe4S4 when reference is made specifically to the core of clusters in proteins, or as indicated above, when the terminal ligands are to be specified.]
During the past decade many iron proteins have been isolated and characterized that have electron-transfer properties but are neither iron-sulfur nor iron-heme proteins. Known examples are:
(a) the lipoxygenases (EC 1.13.11.12);Often these proteins are divided into two classes, i.e. those containing mononuclear iron sites, such as in the lipoxygenase, and those containing dinuclear iron sites, such as in utoferrin and in purple acid phosphatase (ref 30), which are believed to be similar. In these last proteins the two iron sites are assumed to contain the bridge [Fe(O)(RCO2)22Fe], as based upon spectroscopy and comparison with synthetic analogues. However, other bridges, like a phosphate, cannot be excluded.(b) tyrosine 3-monooxygenase (EC 1.14.16.2);
(c) purple acid phosphatase (EC: see Note 1);
(d) uteroferrin (EC: see Note 2);
(e) catechol 2,3-dioxygenase (EC 1.13.11.2);
(f) mandelate 4-monooxygenase (EC 1.14.16.6);
(g) methane monooxygenase (EC 1.14.13.25);
(h) anthranilate 3-monooxygenase (deaminating); from Aspergillus niger; (EC 1.14.13.35).
Note 1 Not yet listed in Enzyme Nomenclature (ref 1).
Note 2 Not yet listed in Enzyme Nomenclature (ref 1).
1. International Union of Biochemistry (1984). Enzyme Nomenclature, Recommendations 1984, Academic Press, Orlando, Florida, U.S.A.
7. International Union of Biochemistry (1979) Enzyme Nomenclature, Recommendations 1978, Academic Press, New York.
16. Nomenclature Committee Of IUB (1978) Nomenclature of Iron-sulfur Proteins, Recommendations 1978, Arch. Biochem. Biophys. 195, 607-610 (1979); Biochem. J. 181, 513-516 (1979); Eur. J. Biochem. 93, 427-430 (1979); also pp. 601-606 of ref. 7. Correction: Eur. J. Biochem. 102, 315-316 (1979).
17. International Union of Pure and Applied Chemistry (1970) Nomenclature of Inorganic Chemistry 2nd edn Butterworths, London 3rd edn (1990), Blackwell, Oxford.
18. Carter, C.W.,Jr., Kraut, J., Freer, S.T., Alden, R.A., Sieker, L.C., Adman, E. & Jensen, L.H. (1972) Proc. Natl Acad. Sci. U.S.A. 69, 3526-3529.
19. Carter, C.W.,Jr., Kraut, J., Freer, S.T. & Alden, R.A. (1974) J. Biol. Chem. 249, 6339-6347.
20. Herskovitz, T., Averill, B.A., Holm, R.H., Ibers, J.A., Phillips, W.D. & Weiher, J.F. (1972) Proc. Natl Acad. Sci. U.S.A. 69, 2437-2441.
21. Bartsch, R.G. (1963) in Bacterial Photosynthesis (Gest, H., San Pietro, A. & Vernon, L.P., ed.) p. 315, Antioch Press, Yellow Springs, Ohio.
22. Sweeny, W. V., Rabinowitz, J. C. & Yoch, D.C. (1975) J. Biol. Chem. 250, 7842-7847.
23. Leigh, J.S.,Jr. & Erecinska, M. (1975) Biochim. Biophys. Acta 387, 95-106.
24. Beinert, H., Emptage, M.H., Dreyer, J.-L., Scott, R.A., Hahn, J.E., Hodgson, K.O. & Thomson, A.J. (1983) Proc. Natl Acad. Sci. U.S.A. 80, 393-396.
25. Hagen, K.S., Watson, A.D. & Holm, R.H. (1983) J. Am. Chem. Soc. 105, 3905-3913.
26. Ghosh, D., O'Donnel, S., Furey, W., Robbins, R.H. & Stout, C.D. (1982) J. Mol. Biol. 158, 73-109.
27. Johnson, M.K., Morningstar, J.E., Cecchini, G. & Ackrell, B.A.C. (1985) Biochem. Biophys. Res. Commun. 131, 653-658.
28. Coucouvanis, D., Salifoglu, M.G., Kanatzidis, M.G., Simopoulos, A. & Papaefthymiou, V. (1984) J. Am. Chem. Soc. 106, 6081-6082.
29. Fee, J.A., Findling, K.L., Yoshida, T., Hille, R., Tarr, G.E., Hearshen, D.O., Dunham, W.R., Day, E.P., Kent, T. A. & Münck, E. (1984) J. Biol. Chem. 259, 124-133.
30. Antanaitis, B.C. &, Aisen, P. (1983) Adv. Inorg. Biochem. 5, 111-134